Rabbit Anti-phospho-p53BP1 (Ser25 + Ser29)antibody
53BP1 (phospho S25); p-53BP1 (phospho S25) ;53BP1 (Phospho-Ser25/29); 53BP1 (Phospho-S25/S29); phospho-p53BP1(Ser25/Ser29); P-53BP1 (Ser25/29); 53 BP1; FLJ41424; MGC138366; p202; p53 binding protein 1; p53 BP1; p53BP1; TP53 BP1; TP53BP1; Tumor protein 53
View History [Clear]
Details
Product Name phospho-p53BP1 (Ser25 + Ser29) Chinese Name 磷酸化p53Binding protein1抗体 Alias 53BP1 (phospho S25); p-53BP1 (phospho S25) ;53BP1 (Phospho-Ser25/29); 53BP1 (Phospho-S25/S29); phospho-p53BP1(Ser25/Ser29); P-53BP1 (Ser25/29); 53 BP1; FLJ41424; MGC138366; p202; p53 binding protein 1; p53 BP1; p53BP1; TP53 BP1; TP53BP1; Tumor protein 53 binding protein 1; Tumor protein p53 binding protein 1; Tumor suppressor p53 binding protein 1; TP53B_HUMAN; tumor protein 53-binding protein, 1. literatures Product Type Phosphorylated anti Research Area Tumour Cell biology Chromatin and nuclear signals Signal transduction Kinases and Phosphatases Binding protein Epigenetics Immunogen Species Rabbit Clonality Polyclonal React Species Mouse, Rat, (predicted: Human, Dog, Pig, Cow, Horse, Rabbit, ) Applications ELISA=1:5000-10000 IHC-P=1:100-500 IHC-F=1:100-500 ICC=1:100-500 IF=1:100-500 (Paraffin sections need antigen repair)
not yet tested in other applications.
optimal dilutions/concentrations should be determined by the end user.Theoretical molecular weight 213kDa Cellular localization The nucleus Form Liquid Concentration 1mg/ml immunogen KLH conjugated Synthesised phosphopeptide derived from human p53BP1 around the phosphorylation site of Ser25/29: ED(p-S)QPE(p-S)QV Lsotype IgG Purification affinity purified by Protein A Buffer Solution 0.01M TBS(pH7.4) with 1% BSA, 0.03% Proclin300 and 50% Glycerol. Storage Shipped at 4℃. Store at -20 °C for one year. Avoid repeated freeze/thaw cycles. Attention This product as supplied is intended for research use only, not for use in human, therapeutic or diagnostic applications. PubMed PubMed Product Detail p53 binding protein 1 (53BP1) plays a critical role in tumor suppression and is a putative substrate of ATM kinase. Upon DNA damage, it is phosphorylated and relocalizes to the presumptive sites of damage, specifically, double strand breaks. This also suggests a role in DNA repair, maintaining genomic stability.
Function:
May have a role in checkpoint signaling during mitosis. Enhances TP53-mediated transcriptional activation. Plays a role in the response to DNA damage.
Subunit:
Interacts with IFI202A. Binds to the central domain of p53/TP53. May form homooligomers. Interacts with DCLRE1C. Interacts with histone H2AFX and this requires phosphorylation of H2AFX on 'Ser-139'. Interacts with histone H4 that has been dimethylated at 'Lys-20' (H4K20me2). Has low affinity for histone H4 containing monomethylated 'Lys-20' (H4K20me1). Does not bind histone H4 containing unmethylated or trimethylated 'Lys-20' (H4K20me3). Has low affinity for histone H3 that has been dimethylated on 'Lys-79'. Has very low affinity for histone H3 that has been monomethylated on 'Lys-79' (in vitro). Does not bind unmethylated histone H3. Interacts with MUM1/EXPAND1. Interacts with CHEK2; modulates CHEK2 phosphorylation at 'Thr-68' in response to infrared. Interacts with MSL1; this interaction may be required for MSL1 DNA repair activity, but not for histone acetyltransferase activity.
Subcellular Location:
Nucleus. Chromosome; centromere; kinetochore. Associated with kinetochores. Both nuclear and cytoplasmic in some cells. Recruited to sites of DNA damage, such as double stand breaks. Methylation of histone H4 at 'Lys-20' is required for efficient localization to double strand breaks.
Post-translational modifications:
Asymmetrically dimethylated on Arg residues by PRMT1. Methylation is required for DNA binding.
Phosphorylated at basal level in the absence of DNA damage. Hyper-phosphorylated in an ATM-dependent manner in response to DNA damage induced by ionizing radiation. Hyper-phosphorylated in an ATR-dependent manner in response to DNA damage induced by UV irradiation.
DISEASE:
Note=A chromosomal aberration involving TP53BP1 is found in a form of myeloproliferative disorder chronic with eosinophilia. Translocation t(5;15)(q33;q22) with PDGFRB creating a TP53BP1-PDGFRB fusion protein.
Similarity:
Contains 2 BRCT domains.
SWISS:
Q12888
Gene ID:
7158
Database links:Entrez Gene: 7158 Human
Entrez Gene: 27223 Mouse
Omim: 605230 Human
SwissProt: Q12888 Human
SwissProt: P70399 Mouse
Unigene: 440968 Human
Unigene: 383499 Mouse
Unigene: 481841 Mouse
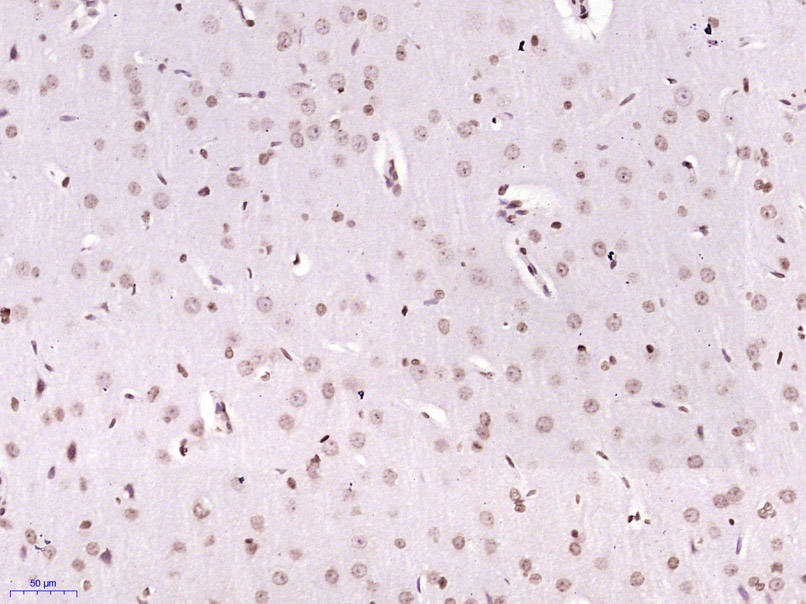
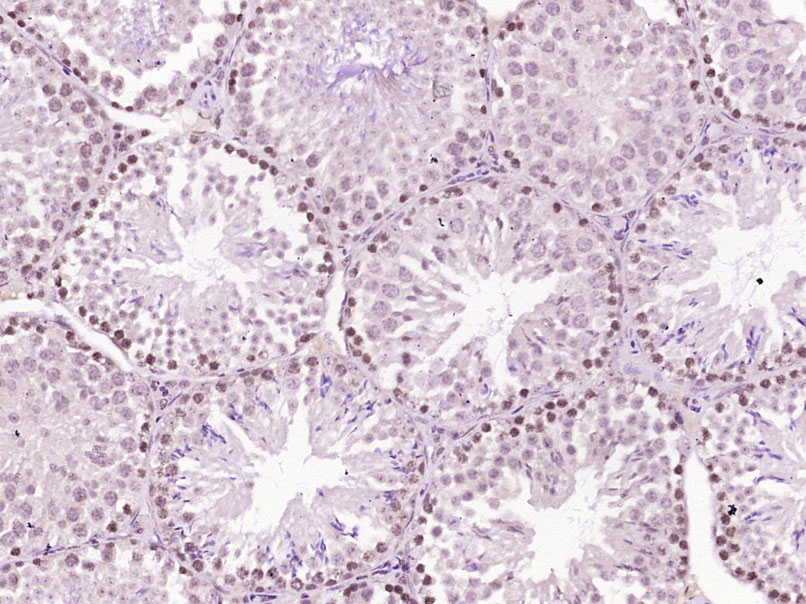
p53Binding protein1(53BP1)是细胞周期检查点激酶蛋白,是新近发现的DNA损伤修复通路中两个非常重要的蛋白激酶MDC1和53BP1蛋白之一,目前多用于Tumour方面的研究。Product Picture
Bought notes(bought amounts latest0)
No one bought this product
User Comment(Total0User Comment Num)
- No comment


 +86 571 56623320
+86 571 56623320
 +86 18668110335
+86 18668110335

